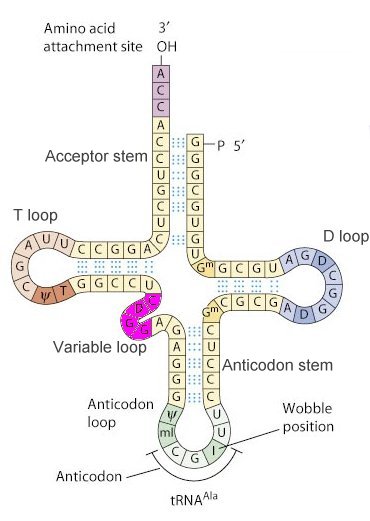
I. IntroductionTransfer RNA (tRNA) serves as the nucleic acid decoding device that reads the triplet genetic code of messenger RNA (mRNA) and causes the insertion of codon-specific amino acids in a growing protein chain during the process of translation in the ribosome. A particular triplet codon in an mRNA is read by a tRNA through its aniticodon loop, which includes a triplet of anticodon residues that base pair with the codon. Each tRNA is charged with a particular amino acid at its 3' end. Although there are 61 codons in the universal genetic code that specify amino acids, most organisms posess fewer than 45 different tRNAs. Some tRNAs can be used to read different codons due to flexibility in the base pairing of the third (3') residue of the mRNA codon and the first (5') residue of the anticodon. These "wobble" base pairs allow for non-Watson-Crick hydorogen bonding, and therefore allow a single tRNA to read multiple codons. The molecule displayed to the left is tRNAPhe that would carry the amino acid phenylalanine attached to its 3' end when appropriately charged by a tRNA synthetase enzyme (see below). II. Structural FeaturesThe secondary structure of a typical tRNA, in this case tRNAAla, is shown in Figure 1, below. The structure consists of hydrogen bonded stems and associated loops, which often contain nucleotides with modified bases (e.g. inosine[I], ribothymidine [T], pseudouridine[Ψ], methylguanosine[D]).
Figure
1. Schematic of tRNA (tRNAAlanine) secondary structure.
The tertiary structure of all tRNAs is similar to that of tRNAPhe, at left, a canonical "L-shaped" molecule. As can be seen, the "cloverleaf" secondary structure shown in Figure 1 results in a complex three dimensional folding of the molecule. The amino acid attachment site at the 3' end and the anticodon loop are observed at the two ends of the "L." The hydrogen bonded stems stabilize the tertiary structure. The Acceptor and Anticodon stems are indicated. Modified bases in the Anticodon, T, and D loops are indicated by thick wireframe. The anticodon bases project from the anticodon loop. The 5' "wobble" base that pairs with the 3' base of the mRNA codon is, in this case, O2'-Methylguanosine.
III. Positioning of tRNAs in the Ribosome and Codon-Anticodon RecognitionShown at left is the crystal structure of the 70S ribosome from Thermus thermophilus, an archaebacterium. Three tRNAs associate with the ribosome in the cavity between the large, 50S and small, 30S subunits (rRNAs are shown in trace, ribosomal proteins in spacefill). Each tRNA is bound in a distinctive site made from structural elements contributed by both ribosomal subunits.The Acceptor stem and 3' end of all three tRNAs are observed to be embedded within the large subunit, whereas the Anticodon stems and loops are found within the small subunit. The A-site tRNA bears an incoming amino acid (not shown) at its 3' end, and the P-site tRNA carries the growing peptide chain (not shown) at its 3' end. Peptide bond formation attaches the peptide to the A-site tRNA's amino acid. The P-site tRNA then moves to the E-site (E stands for "exit"), replacing the former, uncharged E-site tRNA. The A-site tRNA, now bearing the growing peptide, is shifted into the P position. A new tRNA bearing the next amino acid is then brought into the A-site. The A-site tRNA, P-site tRNA, and E-site tRNA exhibit slight conformational differences. However, all adopt the classical "L shape" tertiary structure described above. Note the tight juxtaposition of the 3' Os of the A-site and P-site tRNAs in the peptidyl transferase site of the 50S subunit, providing for peptide bond formation. Let's now turn now to codon-anticodon recognition, which takes place within the small ribosomal subunit. In the structure at left, two phenylalanine codons (UUU) are present in a short mRNA. These codons are are recognized by anticodon bases projecting from the anticodon loops of the A-site and P-site tRNAs. Both tRNAs bear the anticodon residues (AAG) that hydrogen bond with the two UUU codons of the mRNA:
The G-U bonding in third position is an example of the "wobble" base pairing discsussed above. As previously mentioned, wobble pairing allows some codons that differ in the third, 3' base to be recognized by the same tRNA anticodon. This, together with examples of isoaccepting tRNAs that carry the same amino acid but whose anticodons differ in the wobble base, allows for the high degree of degeneracy found in the genetic code. The conformation of the mRNA and the A- and P-site tRNA anticodons helps to ensure that there is no confusion as to which codon should be bound to which tRNA. The conformation is partially achieved by a significant kinking of the mRNA backbone. This results in a distancing of the A- and P-site anticodons. The distance between the 5'G of the P-site anticodon and the 3'A of the A-site anticodon is ~14 Angstroms, much larger than the separation needed if the mRNA were not kinked.
IV. Recognition of tRNA by tRNA SynthetasesAminoacyl-tRNA synthetases play a key role in protein biosynthesis by catalyzing the specific aminoacylation of tRNA at their 3' ends, a two step reaction termed "charging" of tRNA. Class I tRNA synthetases attach an appropriate amino acid to the 2' oxygen of the 3' terminal residue, and class II synthetases attach amino acids at the 3' oxygen. The two steps in the charging reaction are:
It should be clear that in order to ensure proper reading of the genetic code, tRNA charging must match particular anticodon-bearing tRNAs with their correct amino acid. tRNA synthetases must therefore recognize the unique stereochemistry of both individual amino acids and their corresponding tRNAs. The tRNA recognition involves synthetase-anticodon binding as well as other specific interactions, especially at the 3' acceptor stem end of the tRNA. Shown at left is Glutaminyl-tRNA Synthetase bound to tRNAGln. This class I synthetase provides examples of tRNA recognition mechanisms. The synthetase is observed to clasp the tRNA, especially with surface cavities that embrace the anticodon stem and loop and the acceptor stem. With respect to recognition of the anticodon, example protein interactions with the anticodon bases are:
In addition to anticodon bases, numerous other bases interact with the protein. Note especially the interacting bases in the anticodon and acceptor stems. Also, there are obvious electrostatic attractions between positively charged surfaces of the protein and the negatively charged backbone of the tRNA. The active sites of the synthetase, where the two step aminoacyl reaction described above occurs, are quite distant from the anticodon. There are three alpha-beta motifs that are responsible for forming the active site region. ATP and glutamine (not shown) are bound in a pocket formed by the first and second motifs, and the acceptor stem is bound by the third motif. ~38 Angstroms separate ATP from the anticodon. Two short motifs bear residues that are involved in ATP binding. Given the distances involved, how might communication betwen the anticodon recognition domain and the active site domain of the synthetase occur? It has been suggested that two β barrel structures responsible for binding the anticodon stem and loop and which are connected to a long loop might assume a disordered or at least a quite different structure in enzyme which is bound to an incorrect tRNA. The loop is observed to pack against the region containing the ATP binding motifs. Thus, the β barrel structures would align these motifs in the conformation appropriate for catalysis of the step 1 aminoacylation reaction (see above) only when a tRNA with the appropriate anticodon is bound.
V. ReferencesPerona, J.J., Rould, M.A., Steitz, T.A.. Structural basis for transfer RNA aminoacylation by Escheria coli glutaminyl-tRNA synthetase. Biochemistry 32:8758. (1993). Perona, J.J., Rould, M.A., Steitz, T.A.. Structural basis of anticodon loop recognition by glutaminyl-tRNA synthetase. Nature 352:213. (1991). Perona, J.J., Rould, M.A., Steitz, T.A., Soll, D.. Structure of E. coli glutaminyl-tRNA synthetase complexed with tRNA (GLN) and ATP at 2.8A resolution. Science 246:1135. (1989). Shi, H., Moore, P.B.. The crystal structure of yeast phenylalanine tRNA at 1.93 A resolution: a classic structure revisited. RNA 6(8):1091-1105. (2000). Yusupov,
M. M., Yusupova, G. Z., Baucom, A., Lieberman, K., Earnest, T. N.,
Cate, J. H. D., Noller, H. F.: Crystal Structure of the Ribosome at
5.5 A Resolution. Science 292:883-896. (2001).
|
|
Acknowledgement: The format of this web page is modified from a template provided by Dr. Angel Herraez, Bioquimica y Biologia Molecular, Universidad de Alcala,
E-28871, Alcala de Henares (Madrid), Spain.